Difference between revisions of "SUIT-025 O2 mt D057"
From Bioblast
(Created page with "{{MitoPedia |abbr=FNS(Oct,PGM) |description=400px |info=''' ''' - '''SUIT-025''' |application=O2 |SUIT number=D057_...") |
|||
| Line 18: | Line 18: | ||
== Strengths and limitations == | == Strengths and limitations == | ||
::: | :::+ SUIT-025 allows the depletion of endogenous substrates with ADP (1D). | ||
:::+ The protocol provides information on F-pathway in OXPHOS state. The low concentration of malate used in this protocol, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway. | |||
:::+ F-pathway (3Oct-2M.1) can be compared to FN-pathway (5P) in OXPHOS state. | |||
:::+ Pathway control in OXPHOS (F, F(N), FN, FNS and S pathways) can be evaluated. | |||
:::+ Harmonization with many SUIT protocols (up to step 7S). | |||
:::+ Short experimental time. | |||
:::- ET state is not evaluated (for obtaining ET capacity with FNSGp and SGp substrate combinations see [[SUIT-002]]). | |||
:::- | |||
== Compare SUIT protocols == | == Compare SUIT protocols == | ||
Revision as of 17:54, 20 May 2019
Description
Abbreviation: FNS(Oct,PGM)
Reference: - SUIT-025
SUIT number: D057_1D;2M;3Oct;3c;4M;5P;6G;7S;8Rot;9Ama
O2k-Application: O2
- MitoPedia: SUIT -
- SUIT-category: FNS(Oct,PGM)
- SUIT protocol pattern: lineal 1D;2M;3Oct;3c;4M;5P;6G;7S;8Rot-
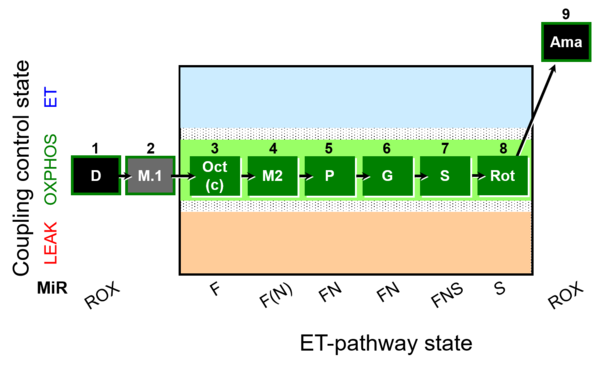
The SUIT-025 protocol is based on SUIT-002 specially designed to give information on F-pathway in OXPHOS state avoiding FAO overestimation in the presence of anaplerotic pathways. Moreover, the pathway control in OXPHOS state (F, F(N), FN, FNS, pathways) can be evaluated by using this SUIT protocol.
Communicated by Doerrier C (last update 2019-05-14)
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1D | ROX | 1D
| ||
| 2M.1 | ||||
| 3Oct | OctMP | F | FAO | 1D;2M.1;3Oct
|
| 3c | OctMcP | F | FAO | 1D;2M.1;3Oct;3c
|
| 4M2 | OctMP | F(N) | FAO | 1D;2M.1;3Oct;4M2
|
| 5P | OctPMP | FN | FAO&CI | 1D;2M.1;3Oct;4M2;5P
|
| 6G | OctPGMP | FN | FAO&CI | 1D;2M.1;3Oct;4M2;5P;6G
|
| 7S | OctPGMSP | FNS | FAO&CI&II | 1D;2M.1;3Oct;4M2;5P;6G;7S
|
| 8Rot | SP | S | CII | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Rot
|
| 9Ama | ROX | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Rot;9Ama
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- + SUIT-025 allows the depletion of endogenous substrates with ADP (1D).
- + The protocol provides information on F-pathway in OXPHOS state. The low concentration of malate used in this protocol, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway.
- + F-pathway (3Oct-2M.1) can be compared to FN-pathway (5P) in OXPHOS state.
- + Pathway control in OXPHOS (F, F(N), FN, FNS and S pathways) can be evaluated.
- + Harmonization with many SUIT protocols (up to step 7S).
- + Short experimental time.
- - ET state is not evaluated (for obtaining ET capacity with FNSGp and SGp substrate combinations see SUIT-002).
Compare SUIT protocols
- SUIT-002 (the reference protocol 2) differs from SUIT-025 in the coupling and pathway control states. SUIT-002 allows the evaluation of pathway control in OXPHOS state (F, F(N), FN, FNS, FNSGp pathways) and in ET state (FNSGp and SGp) whereas in SUIT-025 only F, F(N), FN and FNS pathways in OXPHOS state can be evaluated.
References
MitoPedia concepts: SUIT protocol, SUIT A, Find
MitoPedia methods:
Respirometry