SUIT-001 O2 mt D001: Difference between revisions
Beno Marija (talk | contribs) No edit summary |
No edit summary |
||
| Line 37: | Line 37: | ||
== SUIT O2 DLP for SUIT 1 (RP1) (DatLab 7. | == SUIT O2 DLP for SUIT 1 (RP1) (DatLab 7.3) == | ||
''Updating this page (2018-06-13) for DatLab 7.3'' | |||
:::: [[DL-Protocols]] (DLP) for substrate-uncoupler-inhibitor titrations (see [[MitoPedia: SUIT]]) guide through a sequence of coupling and Electron transfer-pathway states. A specific SUIT protocol can be assigned to O2k-Chamber A or B or both. DL-Protocols for different SUIT protocols can be found in the library of protocols [[MitoPedia: SUIT]]. Instrumental DL-protocols are displayed in the [[DL-Protocols#DL-Protocol_library| DL-Protocol library]]. | :::: [[DL-Protocols]] (DLP) for substrate-uncoupler-inhibitor titrations (see [[MitoPedia: SUIT]]) guide through a sequence of coupling and Electron transfer-pathway states. A specific SUIT protocol can be assigned to O2k-Chamber A or B or both. DL-Protocols for different SUIT protocols can be found in the library of protocols [[MitoPedia: SUIT]]. Instrumental DL-protocols are displayed in the [[DL-Protocols#DL-Protocol_library| DL-Protocol library]]. | ||
Revision as of 13:37, 13 June 2018
Description
Abbreviation: FNSGp(PGM)
Reference: A: SUIT 1 is the RP1 - MiPNet21.06 SUIT RP
SUIT number: 1
MitoPedia concepts:
SUIT protocol,
SUIT A
- MitoPedia: SUIT - SUIT reference protocol RP1
- SUIT-category: FNSGp(PGM)
- SUIT protocol pattern: orthogonal
Harmonized SUIT protocols
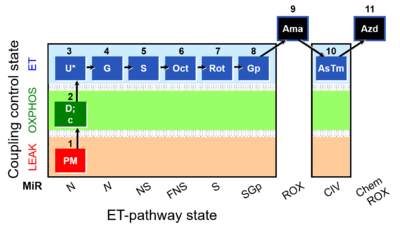
- SUIT RP1:1PM;2D:2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10AsTm;11Azd
- SUIT RP2:1D;2M.1;3Oct;3c;4M2;5P;6G;7S;8Gp;9U;10Rot;11Ama;12AsTm;13Azd
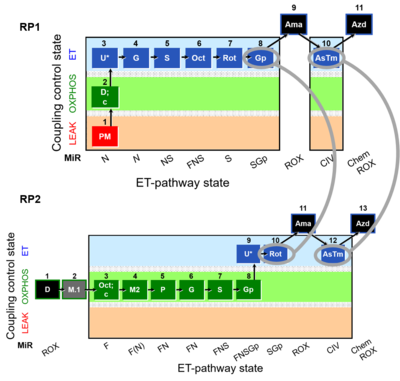 Harmonization between RP1 and RP2
Harmonization between RP1 and RP2
RP1 can be assessed in cells after the selective plasma membrane permeabilization. Digitonin is a mild detergent that permeabilizes plasma membranes. The optimum effective digitonin concentration for complete plasma membrane permeabilization of cultured cells can be determined directly in a respirometric protocol (see Digitonin). The use of intact cells allows the possibility to evaluate respiration in two different modules: intact cells and permeabilized cells. ROUTINE respiration (ce1) is obtained before digitonin is employed, whereas after the use of digitonin we can address differentet Electron transfer-pathway states.
- SUIT RP1 in cells:ce1;1Dig;1PM;2D:2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10Tm;11Azd
- SUIT RP2 in cells:ce1;1Dig;1D;2M.1;3Oct;3c;4M2;5P;6G;7S;8Gp;9U;10Rot;11Ama;12AsTm;13Azdd
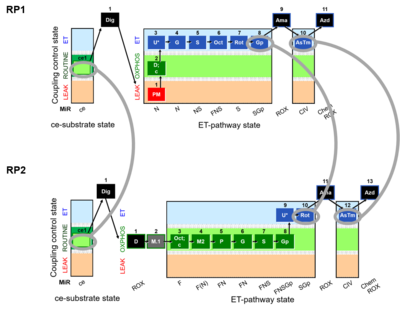 Harmonization between RP1 and RP2 in cells
Harmonization between RP1 and RP2 in cells
In PBMC and PLT the reference protocol RP1 is a different SUIT protocol (absense of Oct): SUIT 3
SUIT O2 DLP for SUIT 1 (RP1) (DatLab 7.3)
Updating this page (2018-06-13) for DatLab 7.3
- DL-Protocols (DLP) for substrate-uncoupler-inhibitor titrations (see MitoPedia: SUIT) guide through a sequence of coupling and Electron transfer-pathway states. A specific SUIT protocol can be assigned to O2k-Chamber A or B or both. DL-Protocols for different SUIT protocols can be found in the library of protocols MitoPedia: SUIT. Instrumental DL-protocols are displayed in the DL-Protocol library.
Download instructions.
To start the download click on theicon. Depending on your browser you will encounter different download scenarios.
(i) A window opens to select whether to "open file with" or to "save file". Please always choose the option "save file"; opening the file directly will not work due to safety reasons.
(ii) When you click the download icon, some browsers will display the text/content of the .DLP file instead of downloading it. In this case go back to this download page and right-click the download icon and choose "save linked file as" from the context-menu. A window will open to save the file. In some cases the suffix of the file might be wrong (e.g. ".HTML") - you must correct the suffix of the file to ".DLP".