SUIT-002 O2 ce-pce D007: Difference between revisions
From Bioblast
No edit summary |
No edit summary |
||
| Line 36: | Line 36: | ||
:::+ SUIT-002 allows the depletion of endogenous substrates with ADP (1D). | :::+ SUIT-002 allows the depletion of endogenous substrates with ADP (1D). | ||
:::+ The protocol provides information on F-pathway in OXPHOS state. The low concentration of malate used in this protocol, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway. | :::+ The protocol provides information on F-pathway in OXPHOS state. The low concentration of malate used in this protocol, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway. | ||
:::+ F-pathway (3Oct-2M.1) compared to FN-pathway (5P) in OXPHOS state. | :::+ F-pathway (3Oct-2M.1) can be compared to FN-pathway (5P) in OXPHOS state. | ||
:::+ Pathway control in OXPHOS (F, F(N), FN, FNS, FNSGp pathways) and in ET state (FNSGp and SGp). | :::+ Pathway control in OXPHOS (F, F(N), FN, FNS, FNSGp pathways) and in ET state (FNSGp and SGp) can be observed. | ||
:::+ Harmonization with [[SUIT-001]] allows to perform both SUIT protocols in parallel. The cross-linked respiratory states can be | :::+ Harmonization with [[SUIT-001]] allows to perform both SUIT protocols in parallel. The cross-linked respiratory states can be statistically used as repeated measurements. | ||
:::+ Harmonization with many SUIT protocols (up to step 7S). | :::+ Harmonization with many SUIT protocols (up to step 7S). | ||
:::+ In SUIT-002, the full set of pathways converging into Q (FNSGp) is obtained in OXPHOS and ET states. Therefore, ''P''/''E'' (8Gp/9U) at high ET capacity can be calculated. | :::+ In SUIT-002, the full set of pathways converging into Q (FNSGp) is obtained in OXPHOS and ET states. Therefore, ''P''/''E'' (8Gp/9U) at high ET capacity can be calculated. | ||
Revision as of 11:08, 19 February 2019
Description
Abbreviation: FNSGp(Oct,PGM)
Reference: A Reference protocol RP2; pce: permeabilized cells - SUIT-002
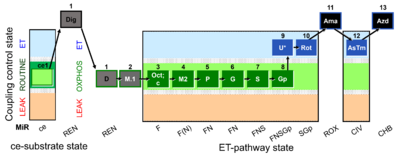
SUIT number: D007_ce1;1Dig;1D;2M;3Oct;3c;4M;5P;6G;7S;8Gp;9U;10Rot;11Ama;12AsTm;13Azd
O2k-Application: O2
- MitoPedia: SUIT - SUIT reference protocol RP2 for permeabilized cells
- SUIT-category: FNSGp(Oct,PGM)
- SUIT protocol pattern: diametral 1D;2M.1;3Oct;3c;4M2;5P;6G;7S;8Gp;9U;10Rot-
References
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| ce1 | ROUTINE | ce1
| ||
| 1Dig | REN | ce1;1Dig
|
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1D | REN | 1D
| ||
| 2M.1 | ||||
| 3Oct | OctMP | F | FAO | 1D;2M.1;3Oct
|
| 3c | OctMcP | F | FAO | 1D;2M.1;3Oct;3c
|
| 4M2 | OctMP | F(N) | FAO | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Gp;9U;10Rot
|
| 5P | OctPMP | FN | F&CI | 1D;2M.1;3Oct;4M2;5P
|
| 6G | OctPGMP | FN | F&CI | 1D;2M.1;3Oct;4M2;5P;6G
|
| 7S | OctPGMSP | FNS | F&CI&II | 1D;2M.1;3Oct;4M2;5P;6G;7S
|
| 8Gp | OctPGMSGpP | FNSGp | F&CI&II&GpDH | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Gp
|
| 9U | OctPGMSGpE | FNSGp | F&CI&II&GpDH | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Gp;9U
|
| 10Rot | SGpE | SGp | CII&GpDH | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Gp;9U;10Rot
|
| 11Ama | ROX | 1D;2M.1;3Oct;4M2;5P;6G;7S;8Gp;9U;10Rot;11Ama
|
| Step | Respiratory state | Pathway control | ET-Complex | Comment |
|---|---|---|---|---|
| ## AsTm | AsTmE | CIV | CIV | |
| ## Azd | CHB |
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- SUIT-002 in combination with SUIT-001 provides a common reference for comparison of respiratory control in a large variety of species, tissues and cell types. Both SUIT protocols provide a mitochondrial mapping which allows:
- 1. to obtain reference values.
- 2. to evaluate mitochondrial physiological diversity, generating a mt-database on comparative mitochondrial physiology.
- 3. to screen specific defects.
- SUIT-001 and SUIT-002 are used in the MitoFit Proficiency test for inter-individual and inter-laboratory reproducibility quality control.
- A succinate concentration of >10 mM may be required for saturating SE capacity.
- + SUIT-002 allows the depletion of endogenous substrates with ADP (1D).
- + The protocol provides information on F-pathway in OXPHOS state. The low concentration of malate used in this protocol, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway.
- + F-pathway (3Oct-2M.1) can be compared to FN-pathway (5P) in OXPHOS state.
- + Pathway control in OXPHOS (F, F(N), FN, FNS, FNSGp pathways) and in ET state (FNSGp and SGp) can be observed.
- + Harmonization with SUIT-001 allows to perform both SUIT protocols in parallel. The cross-linked respiratory states can be statistically used as repeated measurements.
- + Harmonization with many SUIT protocols (up to step 7S).
- + In SUIT-002, the full set of pathways converging into Q (FNSGp) is obtained in OXPHOS and ET states. Therefore, P/E (8Gp/9U) at high ET capacity can be calculated.
- + This protocol can be extended with the Complex IV module.
- - S-pathway in ET state is not obtained (it is obtained in SUIT-001).
- - Lengthy duration of the experiment.
Compare SUIT protocols
- SUIT-002 O2 pce D007 (general SUIT-002 for permeabilized cells) differs from SUIT-002 O2 pce D007a (SUIT-002 for permeabilized PBMCs and PLTs) in the concentration and volume of the titrations.
- SUIT-001 O2 pce D003 (RP1): Harmonized SUIT protocol for permeabilized cells.
MitoPedia concepts:
SUIT protocol,
SUIT A,
Find
MitoPedia methods:
Respirometry
Labels:
SUIT-002