Talk:Fasching M
O2 channel and measuring chamber
Before each experiment
Perform an air calibration. File the obtained values by using the template "O2k-calibration.xlsx", as described in MiPNet19.18 O2k-calibration. Trends over time (e.g. in the zero current R1) can thus be evaluated , and possible irregularities of sensor performance are quickly recognized for intervention (POS service). Close the chamber, do a stirrer test and observe the Flux at closed chamber near to air saturation before introducing the biological sample into the chambers.
After any changes to the system or every few weeks
To do a quick quality control or whenever you are in doubt about the instrumental performance do an Oxygen sensor test.
Record the instrumental background oxygen flux. Beside supplying the required parameters for instrumental background correction the instrumental background experiment is also an excellent quality control experiment for the instrument. When done properly (that is including a stirrer test) all the information obtained in the sensor test is also obtained by recording the background oxygen flux.
Flux at closed chamber near to air saturation
The oxygen flux in a closed chamber at air-calibrated oxygen concentration is an important control parameter. It reflects the consumption of oxygen by the polarographic oxygen sensor (POS). The theoretical value is calculated by DatLab in the O2 Calibration window (Details tab/ Oxygen consumption by POS), see Supplement C in MiPNet06.03 POS-calibration-SOP. The theoretical value at 37°C (O2 slope uncorrected) is usually between 2 and 3 pmol.s-1.ml-1. The actual values should be within +/- of 1 pmol /(s ml). Values higher than 4-5 pmol.s-1.ml-1 at 37° may therefore indicate a biological contamination in the chamber or in the medium (but double check the theoretical value from the Details tab). Lower values may indicate:
- a higher chamber volume: check and if necessary repeat chamber volume calibration
- air bubbles: remove air bubbles
- if a multi-sensor stopper with multiple bores is used: liquid on top of the stopper creating an exchange of liquid between the chamber volume and the outside: aspirate excess liquid from top of stopper
For different fluxes in the left and right chamber (with or without sample) see also Different O2 fluxes in left and right chamber
Raw signal at air saturation
The voltage recorded at air saturation and with an open chamber is an important instrumental quality parameter. It is one result of a full Oxygen sensor test. To access just this parameter observe the Raw signal.
- Fill the chamber with water or medium
- Establish an open chamber (or when using a closed chamber be sure that the O2 concentration is still very near to atmospheric saturation)
- Wait for stabilization
The recorded value will depend on the gain.
Typical recorded voltages at air saturation, in pure water or experimental media, should be
- at Gain 8: off scale ( “9.99V”)
- at Gain 4 : ca. 8 - 10 V (do not use at air saturation, especially if your lab is situated at or near sea level)
- at Gain 2: ca. 4 - 5 V
- at Gain 1: ca. 2 V
The actual value of the raw signal at air saturation (during calibration) depends on, the medium, experimental temperature, barometric pressure, and the sensitivity of the individual POS. At or near sea level higher values are recorded than at higher altitudes. The raw signal is converted to oxygen concentration [µM or nmol/ml] or partial oxygen pressure [kPa] by calibration of the POS [MiPNet12.08]. Each gain requires a separate calibration of the oxygen signal. Therefore, after changing the gain you should not only perform a calibration at air saturation (as always before an experiment), but also a new zero oxygen calibration.
If the recorded value is too high first of all the zero current should be checked. A high zero current will also give a corresponding higher voltage at air saturation. E.g. If a sensor produced a zero current of 1 Volt at gain 2, the voltage at air saturation will typically be 5 to 6 V instead of 4 to 5 V. Please see the entries under zero current for acceptable values of this parameter.
Whether or not a high zero current is found the problem has to be further located by switching components. Typically the problem will be located either on the POS or on the POS connector.
If the zero current if ok, consider the following potential reasons:
- The stirrer speed (rpm) may have been changed, or the stirrer may have been left in the off mode. - Re-set the stirrer speed and re-calibrate.
- The gain setting may have been changed. - Compare calibration values at identical gain settings.
- The block temperature may have been changed. - Compare calibration values at identical temperatures.
- R1 is similar, but the oxygen calibration factor, Fc, may have changed, due to the use of a different oxygen solubility factor of the medium, FM. - Compare calibration factors, Fc, at identical values of FM.
- R1 has changed, but the oxygen calibration factor, Fc, remains similar, if the barometric pressure has changed (e.g. experiments carried out at different altitude). - Compare calibration values, R1, at identical barometric pressure.
The problem occurred after application of a new membrane:
- Lower R1: The membrane thickness may have changed, possibly by inadvertently using two superimposed membranes.
- Lower R1: The membrane may have wrinkles in the area of the cathode.
- Lower R1: The black butyl rubber gasket may have not been applied properly in the centre, now partially blocking the cathode.
- Higher R1: The membrane may have been overstretched, or a thinner membrane is used.
The problem occurred after re-assembly of the O2k chamber:
- Lower R1: The chamber may have been improperly positioned, with the blue POS holder (on the left or right side of the copper block) screwed too far into the chamber housing. This would allow only partial contact between the stirred medium and the cathode. - Follow the O2k-Start instructions (MiPNet12.06, Section 3.1) to insert the chamber properly.
- The electrolyte in the POS may have partially dried out. - Apply a new membrane, following POS-Service instructions (MiPNet08.04).
Reanimating an old oxygraph
Sometimes an oxygraph has not been used for several years. Sometimes just a "start over" is required for a new user. In both cases the following steps may be helpful:
- Get up to date manuals and protocols from http://www.oroboros.at/
- Consider upgrading your DatLab software. If the oxygraph was purchased within the last two years you are probable eligible to a free Datlab upgrade. You might want to consult Oroboros Instruments whether for your particular field of interest an upgrade is recommended (it always is) or strongly recommended.
- Before staring any work on the oxygraph itself read about Electrostatic discharge and how to prevent damage by it.
- Compile an inventory of accessories using the "Start Manual A MiPNet12.06 O2k-Start OROBOROS O2k-Catalogue . Especially for old instruments that were delivered with titanium stoppers you may have to consult your original manual. Spares for instruments with titanium stoppers you can fin in the Catalogue Complete OROBOROS Product List. Order missing accessories.
- Check components that are prone to aging: Are the o-rings on stoppers or chamber holders getting brittle? Do you have spare o-rings? Enough POS membranes?
- Disassemble the chamber assembly reversing the chamber assembly procedures in A MiPNet12.06 O2k-Start. After removing chamber holder, sensor holder and glass chamber clean the exposed copper block, the chamber holder and the sensor holder with plenty of water to dissolve any spilled and tried medium.
- Clean the glass chamber and the stirring bars: MiPNet19.03 O2k-cleaning and ISS
- Clean the sensor connector and its electrical connections: Cleaning the POS connector
- If the sensors (POS) were stored assembled for a very long time they will probable have dried out. Remove membrane holder ring and membrane and wash with water. Make sure that the POS is attached either to the POS senor connector or the blue part of the POS sensor case during this washing (to prevent water from touching the electrical connections of the POS). Perform a full sensor (POS) service MiPNet19.18B_POS-service and mount new membranes to the sensors.
- Reassemble the chamber strictly following A MiPNet12.06 O2k-Start.
- Do a chamber volume calibration A MiPNet12.06 O2k-Start.
- Run the oxygraph with the newly mounted sensors over night
- Perform an Oxygen sensor test
- Consider joining a International oxygraph course
- Please contact [email protected] any time for help. Having done as many of the steps mentioned above will speed up the process. If possible, please attach a DatLab file with a sensor test to your e-mail. (the DatLab file itself not a screen shot). Even if you think your system is running fine you are welcome to send us a sensor test file to check whether the instrumental performance is ok.
DatLab layout does not work
A DatLab layout does not work in the expected way
Question: We have a problem with Datlab 4/5.x. We enter mg as the unit in edit experiment, but when we then chose Layout 06 Specific flux per unit sample it does not change the unit of the flux to per mg - it stays at per ml. How is this possible? Should we reinstall the datlab? is there a bug?
Answer1: One explanation would be that the layout template has been corrupted (saved with wrong settings). You always can re-import (File/Import/DatLab templates) the standard templates from the file "DLTemplates.dlt" available from
- your DatLab install directory in the sub-folder: "DLTemplates"
- your USB stick
- (the most up to date version) DatLab on Oroboros homepage.
Especially, if it is possible to set the correct plot manually (by using "Graph"/"Select Plots" an then selecting "O2flux per mass" for the desired plot, but the "shortcut" via the layout does not work, this must be the reason. Otherwise please send us the DatLab file in which you observe the problem (the DatLab file itself, not a screen shot).
See MiPNet12.07 for a full description how to use the import templates function.
Fasching Mario 10:24, 17 October 2012 (CEST)
Answer 2: A specific possibility for the layout mentioned above (Layout 06 Specific flux per unit sample): The layout will revert to another layout if the amount of sample as set in the "Experiment"/ "Edit" window (mg, cells, units) is zero (otherwise there would be a division by zero). Fasching Mario 17:24, 7 February 2014 (CET)
General guideline for editors and users
Abbreviation for shortcuts
In Mitopedia DatLab the abbreviation field is used for shortcuts. Shortcut keys help provide an easier and quicker method of navigating in DatLab.
Manual as a reference
Further information of each DatLab Mitopedia term can be found in the according manuals, which are listed as a reference.
DatLab error messages
In Mitopedia DatLab a list of common DatLab error messages and the according action/solution are provided. Additionally a link from the corresponding term is added for a quicker access to the appropriate DatLab error message.
DatLab installation support
- Question: DatLab was installed successfully but when trying to run it I get errors complaining about access to some strange looking files.
- Alternative question: When installing DatLab [in this users specific university], it is obligatory to install, and use the O2k, as a PC administrator. Doing this precludes possibility of authorized O2k users operating O2k logged in as guest user (non-administrative account). Is there a workaround to this?
- Answer:
- There are two possible reasons for the described problems:
- The default directory to install DatLab is the "root" directory C:/. Depending on your institutions polices ordinary users may (standard procedure) or may not have write access to this high level directory. So the first fix is to try to install DatLab in a directory in which all users have write access. You can test this by generating and editing a simple text file in this directory.
- It may be that the installation of ALL programs is restricted to administrators by the settings in your institution. Obviously the program has to be installed by an administrator in this case. However, if the administrator installs the program into a directory in which ordinary / guest users have write access, these users will be able to use DatLab without any problems.
- There are two possible reasons for the described problems:
- Note: The following set up does NOT work: The user has READ but not WRITE access to the directory DatLab is installed and wants to save the resulting files in directory he has WRITE access to. Reason: DatLab needs write access in the directory it is installed to to save configurations and temporary files. Solution: Install DatLab to a directory the user has full write access to.
DatLab 5 version history
DatLab 5.2
- Menu [O2k-Multisensor]/[Calibration] The Amp channel can now be calibrated using a multiple point linear regression:
- Select the Amp plot to be calibrated as active plot.
- Mark sections of the experiment for which the concentrations are known.
- Open the calibration window using either [O2k-MultiSensor]/[Calibration] or F5.
- In the left panel the current calibration is shown. The right panel is used to do a new calibration.
- Choose the desired unit for the calibrated signal from the drop down menu beside "Unit".
- Select the marks you want to use for the calibration by ticking the appropriate check boxes in the right panel under the heading „Select marks“. The averaged raw signal values, times, and slopes will be displayed for the selected marks.
- Leave the check boxes beside the slope column unchecked unless calibrating an Amplex Red experiment, see Amplex_red#Calibration_with_H2O2.
- Enter the known concentrations in the column labeled "Conc".
- DatLab will perform a linear regression after each field has been edited. The results, sensitivity and R2, are displayed below the calibration table. You can compare these parameters with those of the currently valid calibration in the left panel.
- Check the quality of the calibration with „Show Graph Plot“. The graph may be exported to the clipboard.
- Apply the new calibration values with „Calibrate“ or continue using the old calibration values with „Cancel“.
Manually entering parameters from a linear regression:
- Choose the desired unit for the calibrated signal from the drop down menu beside "Unit".
- Enter the desired values for sensitivity (slope of a regression signal against concentration) and intercept.
- Apply the Calibration values with „Calibrate“ or continue using the old calibration values with „Cancel“
- Menu [O2k-Multisensor]/[Calibration]: Functions effecting the calibrated signal have been separated from those effecting the slope calculated from the signal by placing the in different tabs: “Calibration” and “Slope”.
- Menu [Experiment]/[Edit]: It now possible to calculate the parameters describing Instrumental background oxygen flux directly in DatLab:
- Perform an instrumental background test and set marks on the O2 slope plot as usually.
- Open [Experiment]/[Edit].
- Click on the “Calculate” button beside the displayed current background correction parameters for the desired chamber.
- Select the marks that should be included in the calculation of background parameters by ticking their check boxes (in a standard background experiment usually all marks will be used).
- Background parameters will be calculated and displayed immediately.
- Control the quality of the regression with “Show graph”.
- As a further quality control you may select to view a graph with the background correction already applied to the slopes. The corrected slopes should be close to zero.
- The graph can be exported using the “Copy to Clipboard” function.
- Close the graph window to return to the linear regression window.
- Apply the background correction by clicking “Save” in the linear regression window and again save in the [Experiment]/[Edit] window.
- The parameters a and b are exported whenever the button "Calibrate and copy to clipboard" is used in the calibration window. Therefore the calibration list template (available at [1]) can be used for quality control and keeping a time track of background parameters.
DatLab 5.1.1.9
- Main menu: [Oxygraph] and [MultiSensor] were integrated into [O2k-MultiSensor]. [O2k-MultiSensor]/ [Calibration] (F5) will always open the Calibration Window for the active plot (O2, pX or Amp).
- pX: [O2k-MultiSensor]/ [Calibration] : Renaming the Amp channel, selecting units for the calibrated signal, setting a factor for the slope calculation, and calculating the slope from raw signal works now for the pX channel analogous to the Amp channel (see above).
- [O2k-MultiSensor]/ [Calibration]: When changing the selected unit for the calibrated signal, DatLab asks whether the calibration should be adjusted accordingly.
- Double clicking "Light on / Light off" in the O2k-Status line will switch on / off the chamber light.
DatLab 5.1.0.130
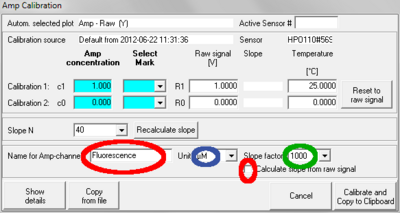
- Menu [MultiSensor]/ [MS Calibration] (Ctrl +F5): For the Amp channel, the user can now set the name of the plot (as it appears e.g. in axis labeling), the unit displayed in the calibrated plot, the multiplication factor used in slope calculation, and whether the slope should be calculated from the calibrated signal or the raw signal. To use these new features:
- Type the desired plot name in the field "Name for Amp-channel". Avoid long names.
- Choose the desired unit for the calibrated signal from the drop down menu beside "Unit". Default: "µM". Changing the unit will not (in DatLab 5.1.0.130) change the calibrated values. Therefore, to avoid confusion, the unit should be chosen before performing a calibration or upgrade to DatLab 5.1.1.7, see below.
- Choose the desired factor for slope calculation from the drop down menu beside "Slope factor". Default: 1000. The correct unit for the slope will be set by DatLab depending on the chosen unit for the calibrated signal and the factor for slope calculation. Changing the factor will recalculate the values for the slope plot correctly.
- To calculate the slope based on the raw signal, activate the checkbox "Calculate from raw signal". If viewing the raw (uncalibrated signal) it is recommended to use this option.
- Click "Calibrate and Copy to Clipboard" to apply all changes.
- Menu [Oxygraph]/ [Stirrer test] (F9) opens a window to enter the desired duration of a Stirrer test (Default: 30 s). After clicking "OK", stirring will be deactivated in both chambers and reactivated after the chosen time.
- Indicator for light in chambers: If the chamber light (not to be confused with light sources for fluorescence) is on, this is indicated in the status line by the text "Light on", highlighted in yellow. If the light is off the text reads "Light off", with no highlighting. This should be helpful when the chamber light is not obviously visible due to Fluorescence-Sensors inserted into the O2k front windows.
- Menu [Oxygraph]/[O2 Calibration] (F5): If the the values for R1 and R0 differ by less than 1 V a warning is displayed. This typically happens when either the same mark is chosen for R1 and R0 or when the two marks for R1 and R0 are set erroneously to the same "state" of the chamber, that is both regions are either for the "open chamber" or for the "closed chamber, no oxygen" state. Check the selected marks when seeing this warning. It is possible to ignore the warning and continue with "Calibrate and Copy to Clipboard".
- Menu [Oxygraph]/ [O2k Control] (F7) / [Configuration]: A field "pX Reference Electrode" was added. A serial number or other designation can be entered.
- Menu [Experiment]/ [View protocol] (Ctrl+F3): The information is now more clearly structured to discern between parameters used during a calibration (may originate from another file, different condition) and the parameters used when the file was recorded. Both, calibration information and the information about actually used parameters were expanded for the pX and Amp channels.
DatLab 5.1.0.20
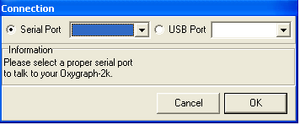
- DatLab 5 supports connecting the oxygraph to a computer via the computers USB port and a standard USB cable (requires O2k-Core or O2k Series E and above). Additionally, DatLab 5 still supports connection to a computer via the RS232 port. Therefore, it is possible and highly recommended to upgrade all oxygraphs in a lab to DatLab 5. When connecting to the O2k for the first time, the type of connection (USB or RS232) is set by radio buttons. When connecting via the USB port any detected oxygraphs will be identified by their serial number and can be selected in the drop down menu to the right of "USB Port". When connecting via the RS232 port the proper serial port "Com..." can be selected in the drop down menu to the right of "Serial Port".
- DatLab 5 allows to open DatLab files by double clicking them in the Windows file manager or similar programs. First time use: Double click any DatLab file in the Windows file manager, select "open this file type always with this program", select "browse", then navigate to the file Datlab5.exe. Usually this file will be in C:\Datlab\DatLab5.exe. Depending on your Windows settings the extension '.exe' may not be displayed: In this case you can differentiate between DatLab5.exe and DatLab5.ini by right clicking and selecting "properties". The exact procedure will depend on the version and language of your Windows operating system. After doing this once, Windows will always use DatLab 5 to open DatLab files.
- Menu [Graph]/[Scaling] (F6): It is possible to set the maximum value for the Y axis of a plot. When changing one or two parameters (minimum, maximum, or range) the remaining parameters will be adjusted automatically after pressing "OK" or clicking in a different field. Fill out only the fields Minimum and Range to simulate the behavior of DatLab 4.
- Menu [Graph]/[Scaling] (F6):It is possible to selectively copy scaling settings to other windows by using the "Apply to Graph 1", Apply to Graph 2", etc buttons. These buttons replace the "Apply to all" button. This will be helpful in MultiSensor applications when e.g. two windows should show oxygen plots (with identical scaling) and two windows should show fluorescence plots (with identical scaling).
- The number of points used for slope calculations can now be changed for the pX and the Amp channels. Select [MultiSensor]/[MS Calibration]: Slope N: Select the number of data points (N = 40 to N = 5 in intervals of 5) used to calculate the slope. A high value of N yields a highly smoothed curve, whereas a low value of N improves time resolution.
- The slopes for the pX and Amp channel are now calculated with the correct sign for the first order time derivative. That is, if the calibrated signal decreases the slope will have a negative sign, if it increases the sign of the corresponding slope will be positive. The slope for the O2 channels is still calculated to give a positive sign for a decreasing signal, signifying a "positive " respiration.
- Proton production rates can be calculated on line. In the menu select [Plots]/[Proton Flux], see O2k-pH ISE-Module.
- The amperometric channel that, due to its main use for "NO electrodes", was labeled "NO" in DatLab 4 was renamed to "Amp" to reflect its versatile nature. It is used e.g. to record the data from the O2k-Fluorescence LED2-Module.
- Faster processing of DatLab Data (this required a larger minimum file size).
DatLab for Apple
Hi Mario, By the way, do you have any version of DataLab that can be installed in Apple Mac system? Or, do anyone from your company know how it get it works?
Hi Peter,
We develop DatLab only for Windows, because its the platform all our customers can live with and most customers have to live with. We are aware that it is not the best platform for scientific software (this might be Linux > Apple > Windows), but unfortunately we do not have a choice here. Even the ongoing software developing for one platform is a very major and cost intensive project for a company of our size, therefore we can not do this for multiple platforms. However, there were rumors that DatLab runs on a Windows emulator software for Apple. This was before we hat this wiki for collection of this kind of user feedback :-( . We do not have experiences in this on our own. I have only experience with running DatLab under a Windows emulator for Linux. This works always for "off line " data analysis and for some DatLab versions even for on-line data accumulation. Without being any sort of Apple expert, I would guess things might be similar for Apple based windows emulators.
Any experiences from customers in this regard would be highly welcome of this page
best greetings Mario --Fasching Mario 09:13, 22 February 2012 (CET)
Hi Mario, By the way, to open DatLab file in Mac system, I install a program called "VitualBox" which is free for basic version. With that, I can install windows in my Mac and therefore all other programs that can be installed in PC. I can open DatLab file without any problem. I am not sure if I can run DatLab to collect data in real-time. Never try it yet since I use other PC to run experiment.
Hi Mario,
we have used an Apple MacBook running DatLab 4 under Parallels 7 for both collecting and analysing data on expedition. It works all right, not superbly, as there are some issues with the right click and a couple of shortcuts. Once you get used to this, you can do everything in DatLab that you can do on a PC. Best wishes
Felix
DatLab response is slow
- Problem: The DatLab software is extremely slow. Opening an experiment can take up to several minutes (usually around 1-2 min) and the same goes for any kind of operation undertaken in the software once the experiment is open (changing axis scale, shifting time frame etc.)
- Solutions: A frequent problem encountered is that DatLab files are saved remotely (on a server) without the user realizing that this is the case. Files that are just being acquired (during the measurement) should always be saved locally. OFF line line files can usually be processed remotely but this depends on the the network speed. To see if the problem is caused by remote files, make sure to save a DatLab file locally on your computer ("C drive") and determine if this does improve performance. For working with DatLab in off-line mode (processing stored files) working via a Network is usually possible, but depending on Network speed and size of file it may sometimes be advisable to copy large files to a local drive (e.g. when working via a WLAN connection).
- There may be an inadvertently small setting for the data sampling interval in DatLab (default 2 s). Very small data sampling intervals are usually necessary only for kinetics experiments. Such very small data sampling intervals may lead to a slow responding DatLab, especially for long recordings. Check the 'Experiment' / 'Edit' window to see the settings for an already recorded file or set the data sampling interval for new files in the 'Oxygraph' / 'O2k Control' window
Export data from DatLab
Export data from DatLab may refer to
- exporting time averaged values for certain regions of the experiment: [Marks] / [Statistics]
- exporting parts of or an entire plot in a point per point fashion to an text file for advanced analysis or drawing the plot outside DatLab: [File]/[Export]/[Data to text File]
- exporting events [File]/[Export]/[Events to text File]
See C MiPNet12.07 DatLab Guide for descriptions of all three functions.
Problems with exporting plots to a text (.csv) file
- Export data does not work for DatLab versions below 4.2.1
- Make sure that DatLab and the Spreadsheet program use the same decimal separator. DatLab takes the character for the decimal separator form the windows system settings.
- It seems that with some recent versions of Excel importing a comma separated text file involves playing around with settings. Solutions:
- (Excel) Set the separator character in the DatLab export function function to the one expected by Excel (may not be possible if Excel expects e.g. a blank).
- (Excel) Change the WINDOWS (not Excel!) settings (Regions and languages / more options) so that the "list separator" character is the same as used by the DatLab export function. See also this link to Microsoft.
- Libre(Open)Office still shows the behavior known from previous Excel versions: double clicking a file opens a preview window and the separator can be chosen interactively.
DatLab scripting function
We do not expect many users to generate scripts on their own (same as with with Graph layouts, TIP setups, etc). But we can generate new scripts without modifying the program itself and make them available to customers e.g. via the Import templates function (or by just copying the file). The functions described here are only available for DatLab 5.1.1.101 and above. The functions may also be removed for released versions of DatLab with a higher version number.
Help
The scripting function to generate new plots is started with [Ctrl+Alt+D], this will open the "calculated columns" window.
A new script is generated by clicking "Generate Plot"
The generation of the scripts themselves is explained in the "help function" of the script window, where also an example script can be found.
A script may be tested e.g. by clicking "compile".
Once a script is finished there are two save buttons:
"Save Macro" (right upper corner) will save the new script to a separate file in the DatLab user directory (in \DatLab\APPDATA\CALC), so it can be reused (and modified) later for other files just like a graph layout, TIP setup et all. In this way a script can also be shared with other users or incorporated to the default functions available for all users.
"Generate plot" or "Update plot" (lower right corner) will save the script to the active DatLab file, that is a new plot is generated. After saving in this way, the new plot can be used: In the Main Menu select [Graph]/[Select Plots]. New plots will be listed at the end of the list. Select the new plots for the desired windows as usual. This script used to generate a new plot is also saved with the particular DatLab file. So if the same DatLab file is opened later, the plot (and its script) will be available, and it is not necessary to load the script form the user directory again.
Help in DatLab
- This text needs to be edited!
Define Plot-Variables by clicking the unit in the list with the right mouse button.
The result of the calculation has to be assigned to the variable "FuncResult". To define additional variables (for easier reading of the script) by the keyword "var" and a type definition ("float"). Each line has to have a trailing semicolon. Example 1 (a little bit oversized to show variable use): var Sum: Float;var Divider: Integer; Sum:=A+B;Divider:=2;FuncResult:=Sum/Divider; Example 2 (same calculation, shortened to one line):FuncResult:=(A+B)/2;
Popular Bioblast page
- Fasching M has been accessed more than
- 15,000 times (2019-12-11)
- Fasching M has been accessed more than